Evolution of Antibiotic Resistance Makes the Case for a Creator
What would it be like to live in a world without antibiotics?
It isn’t that hard to imagine, because antibiotics weren’t readily available for medical use until after World War II. And since that time, widespread availability of antibiotics has revolutionized medicine. However, the ability to practice modern medicine is being threatened because of the rise of antibiotic-resistant bacteria. Currently, there exists a pressing need to understand the evolution of antibiotic-resistant strains and to develop new types of antibiotics. Surprisingly, this worthy pursuit has unwittingly stumbled upon evidence for a Creator’s role in the design of biochemical systems.
Alexander Fleming (1881–1955) discovered the first antibiotic, penicillin, in 1928. But it wasn’t until Ernst Chain, Howard Florey, and Edward Abraham purified penicillin in 1942 and Norman Heatley developed a bulk extraction technique in 1945 that the compound became available for routine medical use.

Figure 1: Alexander Fleming. Image Credit: Wikipedia
Prior to this time, people often died from bacterial infections. Complicating this vulnerability to microbial pathogens was the uncertain outcome of many medical procedures. For example, patients often died after surgery due to complications arising from infections.
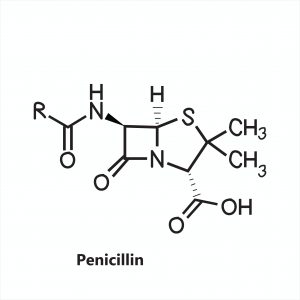
Figure 2: A generalized structure for penicillin antibiotics. Image credit: Shutterstock
Bacterial Resistance Necessitates New Antibiotics
Unfortunately, because of the growing threat of superbugs—antibiotic-resistant strains of bacteria—health experts around the world worry that we soon will enter into a post-antibiotic era in which modern medicine will largely revert to pre-World War II practices. According to Dr. David Livermore, laboratory director at Public Health England, which is responsible for monitoring antibiotic-resistant strains of bacteria, “A lot of modern medicine would become impossible if we lost our ability to treat infections.”1
Without antibiotics, people would routinely die of infections that we easily treat today. Abdominal surgeries would be incredibly risky. Organ transplants and chemotherapy would be out of the question. And the list continues.
The threat of entering into a post-antibiotic age highlights the desperate need to develop new types of antibiotics. It also highlights the need to develop a better understanding of evolutionary processes that lead to the emergence of antibiotic resistance in bacteria.
Recently, a research team from Michigan State University (MSU) published a report that offers insight into the latter concern. These researchers studied the evolution of antibiotic resistance in bacteria that had been serially cultured in the laboratory for multiple decades in media that was free from antibiotics.2 Through this effort, they learned that the genetic history of the bacterial strain plays a key role in its acquisition of resistance to antibiotics.
This work has important implications for public health, but it also carries theological implications. The decades-long experiment provides evidence that the elegant designs characteristic of biochemical and biological systems most likely stem from a Creator’s handiwork.
The Long-Term Evolution Experiment
To gain insight into the role that genetic history plays in the evolution of antibiotic resistance, the MSU researchers piggy-backed on the famous Long-Term Evolution Experiment (LTEE) at Michigan State University. Inaugurated in 1988, the LTEE is designed to monitor evolutionary changes in the bacterium E. coli, with the objective of developing an understanding of the evolutionary process.
Figure 3: A depiction of E. coli. Image Credit: Shutterstock
The LTEE began with a single cell of E. coli that was used to generate twelve genetically identical lines of cells. The twelve clones of the parent E. coli cell were separately inoculated into a minimal growth medium containing low levels of glucose as the only carbon source. After growing overnight, an aliquot (equal fractional part) of each of the twelve cultures was transferred into fresh growth media. This process has been repeated every day for about thirty years. Throughout the experiment, aliquots of cells have been frozen every 500 generations. These frozen cells represent a “fossil record” of sorts that can be thawed out and compared to current and other past generations of cells.
Relaxed Selection and Decay of Antibiotic Resistance
In general, when a population of organisms no longer experiences natural selection for a particular set of traits (antibiotic resistance, in this case), the traits designed to handle that pressure may experience functional decay as a result of mutations and genetic drift. This process is called relaxed selection.
In the case of antibiotic resistance, when the threat of antibiotics is removed from the population (relaxed selection), it seems reasonable to think that antibiotic resistance would decline in the population because in most cases antibiotic resistance comes with a fitness cost. In other words, bacterial strains that acquire antibiotic resistance face a trade-off that makes them less fit in environments without the antibiotic.
Genetic History and the Re-Evolution of Antibiotic Resistance
In light of this expectation, the MSU researchers wondered how readily bacteria that have experienced relaxed selection can overcome loss of antibiotic resistance when the antibiotic is reintroduced to the population.
To explore this question, the researchers examined the evolution of antibiotic resistance in the LTEE ancestor by exposing it to a set of different antibiotics and compared its propensity to acquire antibiotic resistance with four strains of E. coli derived from the LTEE ancestor (that underwent 50,000 generations of daily growth and transfer into fresh media in the absence of exposure to antibiotics).
As expected, the MSU team discovered that 50,000 generations of relaxed selection rendered the four strains more susceptible to four different antibiotics (ampicillin, ceftriaxone, ciprofloxacin, and tetracycline) compared to the LTEE ancestor. When they exposed these strains to the different antibiotics, the researchers discovered that acquisition of antibiotic resistance was idiosyncratic: some strains more readily evolved antibiotic resistance than the LTEE ancestor and others were less evolvable.
Investigators explained this difference by arguing that during the period of relaxed selection some of the strains experienced mutations that constrained the evolution of antibiotic resistance, whereas others experienced mutations that potentiated (activated) the evolution of antibiotic resistance. That is, historical contingency has played a key role in the acquisition of antibiotic resistance. Different bacterial lineages accumulated genetic differences that influence their capacity to evolve and adapt in new directions.
Historical Contingency
This study follows on the heels of previous studies that demonstrate the historical contingency of the evolutionary process.3 In other words, chance governs biological and biochemical evolution at its most fundamental level. As the MSU researchers observed, evolutionary pathways consist of a historical sequence of chance genetic changes operated on by natural selection (or that experience relaxed selection), which, too, consists of chance components.
Because of the historically contingent nature of the evolutionary process, it is highly unlikely that the same biological and biochemical designs should appear repeatedly throughout nature. In his book Wonderful Life, Stephen Jay Gould used the metaphor of “replaying life’s tape.” If one were to push the rewind button, erase life’s history, and then let the tape run again, the results would be completely different each time.4
The “Problem” of Convergence
And yet, we observe the opposite pattern in biology. From an evolutionary perspective, it appears as if the evolutionary process independently and repeatedly arrived at the same outcome, time and time again (convergence). As evolutionary biologists Simon Conway Morris and George McGhee point out in their respective books Life’s Solution and Convergent Evolution, identical evolutionary outcomes are a widespread feature of the biological realm.5
Scientists see these repeated outcomes at ecological, organismal, biochemical, and genetic levels. To illustrate the pervasiveness of convergence at the biochemical level, I describe 100 examples of convergence in my book The Cell’s Design.6
From my perspective, the widespread occurrence of convergent evolution is a feature of biology that evolutionary theory can’t genuinely explain. In fact, given the clear-cut demonstration that the evolutionary process is historically contingent, I see the widespread occurrence of convergence as a failed scientific prediction for the evolutionary paradigm.
Evolution in Bacteria Doesn’t Equate to Large-Scale Evolution
The evolution of E. coli in the LTEE doesn’t necessarily validate the evolutionary paradigm. Just because such change is observed in a microbe doesn’t mean that evolutionary processes can adequately account for life’s origin and history, and the full range of biodiversity.
Convergence and the Case for Creation
Instead of viewing convergent features as having emerged through repeated evolutionary outcomes, we could understand them as reflecting the work of a divine Mind. In this scheme, the repeated origins of biological features equate to the repeated creations by an intelligent Agent who employs a common set of solutions to address a common set of problems facing unrelated organisms.
Sadly, many in the scientific community are hesitant to embrace this perspective because they are resistant to the idea that design and purpose may play a role in biology. But, one can hope that someday the scientific community will be willing to move into a post-evolution future as the evidence for a Creator’s role in biology mounts.
Resources
- The Cell’s Design: How Chemistry Reveals the Creator’s Artistry by Fazale Rana (book)
The Historical Contingency of the Evolutionary Process
- “Inability to Repeat Past Dooms Evolution” by Fazale Rana (article)
- “Historical Contingency and the Improbability of Protein Evolution, Part 1” by Fazale Rana (article)
- “Historical Contingency and the Improbability of Protein Evolution, Part 2” by Fazale Rana (article)
- “Long-Term Evolution Experiment: Evidence for the Evolutionary Paradigm? Part 1” by Fazale Rana (article)
- “Long-Term Evolution Experiment: Evidence for the Evolutionary Paradigm? Part 2” by Fazale Rana (article)
Microbial Evolution and the Validity of the Evolutionary Paradigm
- “Do Plastic-Eating Bacteria Dump the Case for Creation?” by Fazale Rana (article)
- “Does the Evolution of Caffeine-Eating Bacteria Stimulate the Case for Biological Evolution?” by Fazale Rana (article)
Check out more from Reasons to Believe @Reasons.org
Endnotes
- Sarah Bosley, “Are You Ready for a World without Antibiotics?” The Guardian, August 12, 2010, https://www.theguardian.com/society/2010/aug/12/the-end-of-antibiotics-health-infections.
- Kyle J. Card et al., “Historical Contingency in the Evolution of Antibiotic Resistance after Decades of Relaxed Selection,” PLoS Biology 17, no. 10 (October 23, 2019): e3000397, doi:10.1371/journal.pbio.3000397.
- Zachary D. Blount et al., “Historical Contingency and the Evolution of a Key Innovation in an Experimental Population of Escherichia coli,” Proceedings of the National Academy of Sciences USA 105, no. 23 (June 10, 2008): 7899-7906, doi:10.1073/pnas.0803151105.
- Stephen Jay Gould, Wonderful Life: The Burgess Shale and the Nature of History (New York: W.W. Norton & Company, 1990).
- Simon Conway Morris, Life’s Solution: Inevitable Humans in a Lonely Universe (New York: Cambridge University Press, 2003); George McGhee, Convergent Evolution: Limited Forms Most Beautiful (Cambridge, MA: MIT Press, 2011).
- Fazale Rana, The Cell’s Design: How Chemistry Reveal the Creator’s Artistry (Grand Rapids, MI: Baker, 2008).





