Does Evolutionary Bias Create Unhealthy Stereotypes about Pseudogenes?
Truth be told, we all hold to certain stereotypes whether we want to admit it or not. Though unfair, more often than not, these stereotypes cause little real damage.
Yet, there are instances when stereotypes can be harmful—even deadly. As a case in point, researchers have shown that stereotyping disrupts the healthcare received by members of so-called disadvantaged groups, such as African Americans, Latinos, and the poor.1
Healthcare providers are frequently guilty of bias towards underprivileged people. Often, the stereotyping is unconscious and unintentional. Still, this bias compromises the medical care received by people in these ethnic and socioeconomic groups.
Underprivileged patients are also guilty of stereotyping. It is not uncommon for these patients to perceive themselves as the victims of prejudice, even when their healthcare providers are genuinely unbiased. As a result, these patients don’t trust healthcare workers and, consequently, withhold information that is vital for a proper diagnosis.
Fortunately, psychologists have developed best practices that can reduce stereotyping by both healthcare practitioners and patients. Hopefully, by implementing these practices, the impact of stereotyping on the quality of healthcare can be minimized over time.
Recently, a research team from Australia identified another form of stereotyping that holds the potential to negatively impact healthcare outcomes.2 In this case, the impact of this stereotyping isn’t limited to disadvantaged people; it affects all of us.
A Bias Against Pseudogenes
These researchers have uncovered a bias in the way life scientists view the human genome (and the genomes of other organisms). Too often they regard the human genome as a repository of useless, nonfunctional DNA that arises as a vestige of evolutionary history. Because of this view, life scientists and the biomedical research community eschew studying regions of the human genome they deem to be junk DNA. This posture is not unreasonable. It doesn’t make sense to invest precious scientific resources to study nonfunctional DNA.
Many life scientists are unaware of their bias. Unfortunately, this stereotyping hinders scientific advance by delaying discoveries that could be translated into the clinical setting. Quite often, supposed junk DNA has turned out to serve a vital purpose. Failure to recognize this function not only compromises our understanding of genome biology, but also hinders biomedical researchers from identifying defects in these genomic regions that contribute to genetic diseases and disorders.
As psychologists will point out, acknowledging bias is the first step to solving the problems that stereotyping causes. This is precisely what these researchers have done by publishing an article in Nature Review Genetics.3 The team focused on DNA sequence elements called pseudogenes. Traditionally, life scientists have viewed pseudogenes as the remnants of once functional genes. Biologists have identified three categories of pseudogenes: (1) unitary, (2) duplicated, and (3) processed.
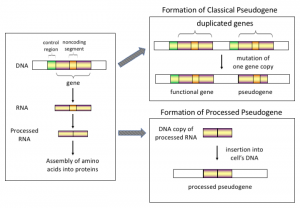
Researchers categorize DNA sequences as pseudogenes based on structural features. Such features indicate to the investigators that these sequence elements were functional genes at one time in evolutionary history, but eventually lost function due to mutations or other biochemical processes, such as reverse transcription and DNA insertion. Once a DNA sequence is labeled a pseudogene, bias sets in and researchers just assume that it lacks function—not because it has been experimentally demonstrated to be nonfunctional, but because of the stereotyping that arises out of the evolutionary paradigm.
The authors of the piece acknowledge that “the annotation of genomics regions as pseudogenes constitutes an etymological signifier that an element has no function and is not a gene. As a result, pseudogene-annotated regions are largely excluded from functional screen and genomic analyses.”4 In other words, the “pseudogene” moniker biases researchers to such a degree that they ignore these sequence elements as they study genome structure and function without ever doing the hard, experimental work to determine whether it is actually nonfunctional.
This approach is clearly misguided and detracts from scientific discovery. As the authors admit, “However, with a growing number of instances of pseudogene-annotated regions later found to exhibit biological function, there is an emerging risk that these regions of the genome are prematurely dismissed as pseudogenic and therefore regarded as void of function.”5
Discovering Function Despite Bias
The harmful effects of this bias become evident as biomedical researchers unexpectedly stumble upon function for pseudogenes, time and time, again, not because of the evolutionary paradigm, but despite it. These authors point out that many processed pseudogenes are transcribed and, of those, many are translated to produce proteins. Many unitary and duplicated pseudogenes are also transcribed. Some are also translated into proteins, but a majority are not. Instead they play a role in gene regulation as described by the competitive endogenous RNA hypothesis.
Still, there are some pseudogenes that aren’t transcribed and, thus, could rightly be deemed nonfunctional. However, the researchers point out that the current experimental approaches for identifying transcribed regions are less than ideal. Many of these methods may fail to detect pseudogene transcripts. However, as the researchers point out, even if a pseudogene isn’t transcribed it still may serve a functional role (e.g., contributing to chromosome three-dimensional structure and stability).
This Nature article raises a number of questions and concerns for me as a biochemist:
- How widespread is this bias?
- If this type of stereotyping exists toward pseudogenes, does it exist for other classes of junk DNA?
- How well do we really understand genome structure and function?
- Do we have the wrong perspective on the genome, one that stultifies scientific advance?
- Does this bias delay the understanding and alleviation of human health concerns?
Is the Evolutionary Paradigm the Wrong Framework to Study Genomes?
Based on this article, I think it is safe to conclude that we really don’t understand the molecular biology of genomes. We are living in the midst of a scientific revolution that is radically changing our view of genome structure and function. The architecture and operations of genomes appear to be far more elegant and sophisticated than anyone ever imagined—at least within the confines of the evolutionary paradigm.
This insight also leads me to question if the evolutionary paradigm is the proper framework for thinking about genome structure and function. From my perspective, treating biological systems as the Creator’s handiwork provides a superior approach to understanding the genome. A creation model approach promotes scientific advance, particularly when the rationale for the structure and function of a particular biological system is not apparent. This expectation forces researchers to keep an open mind and drives further study of seemingly nonfunctional, purposeless systems with the full anticipation that their functional roles will eventually be uncovered.
Over the last several years, I have raised concerns about the bias life scientists have harbored as they have worked to characterize the human genome (and genomes of other organisms). It is gratifying to me to see that there are life scientists who, though committed to the evolutionary paradigm, are beginning to recognize this bias as well.
The first step to addressing the problem of stereotyping—in any sector of society—is to acknowledge that it exists. Often, this step is the hardest one to take. The next step is to put in place structures to help overcome its harmful influence. Could it be that part of the solution to this instance of scientific stereotyping is to grant a creation model approach access to the scientific table?
Resources
- Who Was Adam? A Creation Model Approach to the Origin of Humanity, 2nd exp. ed., by Fazale Rana with Hugh Ross (book)
- The Cell’s Design: How Chemistry Reveals the Creator’s Artistry by Fazale Rana (book)
Pseudogene Function
- “Pseudogene Decoy Reveals Hidden Evidence for Design” by Fazale Rana (article)
- “Functional Pseudogenes Are Everywhere!” by Fazale Rana (article)
- “High Levels of Pseudogene Expression Help Silence the Case for Common Descent” by Fazale Rana (article)
- “Scientific Upset: More Functions for Pseudogenes” by Fazale Rana (article)
- “Pseudogene Discovery Pains Evolutionary Paradigm” by Fazale Rana (article)
- “Unitary Pseudogenes Have Function! Part 1” by Fazale Rana (article)
- “Unitary Pseudogenes Have Function! Part 2” by Fazale Rana (article)
- “Competitive Endogenous RNA Hypothesis Supports the Case for Creation” by Fazale Rana (article)
The Evolutionary Paradigm Hinders Scientific Advance
- “Does the Evolutionary Paradigm Stymie Scientific Advance?” by Fazale Rana (article)
- “Evolution’s Flawed Approach to Science” by Fazale Rana (article)
Check out more from Reasons to Believe @Reasons.org
Endnotes
- For example, see Joshua Aronson et al., “Unhealthy Interactions: The Role of Stereotype Threat in Health Disparities,” American Journal of Public Health 103 (January 1, 2013): 50–56, doi:10.2105/AJPH.2012.300828.
- Seth W. Cheetham, Geoffrey J. Faulkner, and Marcel E. Dinger, “Overcoming Challenges and Dogmas to Understand the Functions of Pseudogenes,” Nature Reviews Genetics 21 (March 2020): 191–201, doi:10.1038/s41576-019-0196-1.
- Cheetham, Faulkner, and Dinger, 191–201.
- Cheetham, Faulkner, and Dinger, 191–201.
- Cheetham, Faulkner, and Dinger, 191–201.





